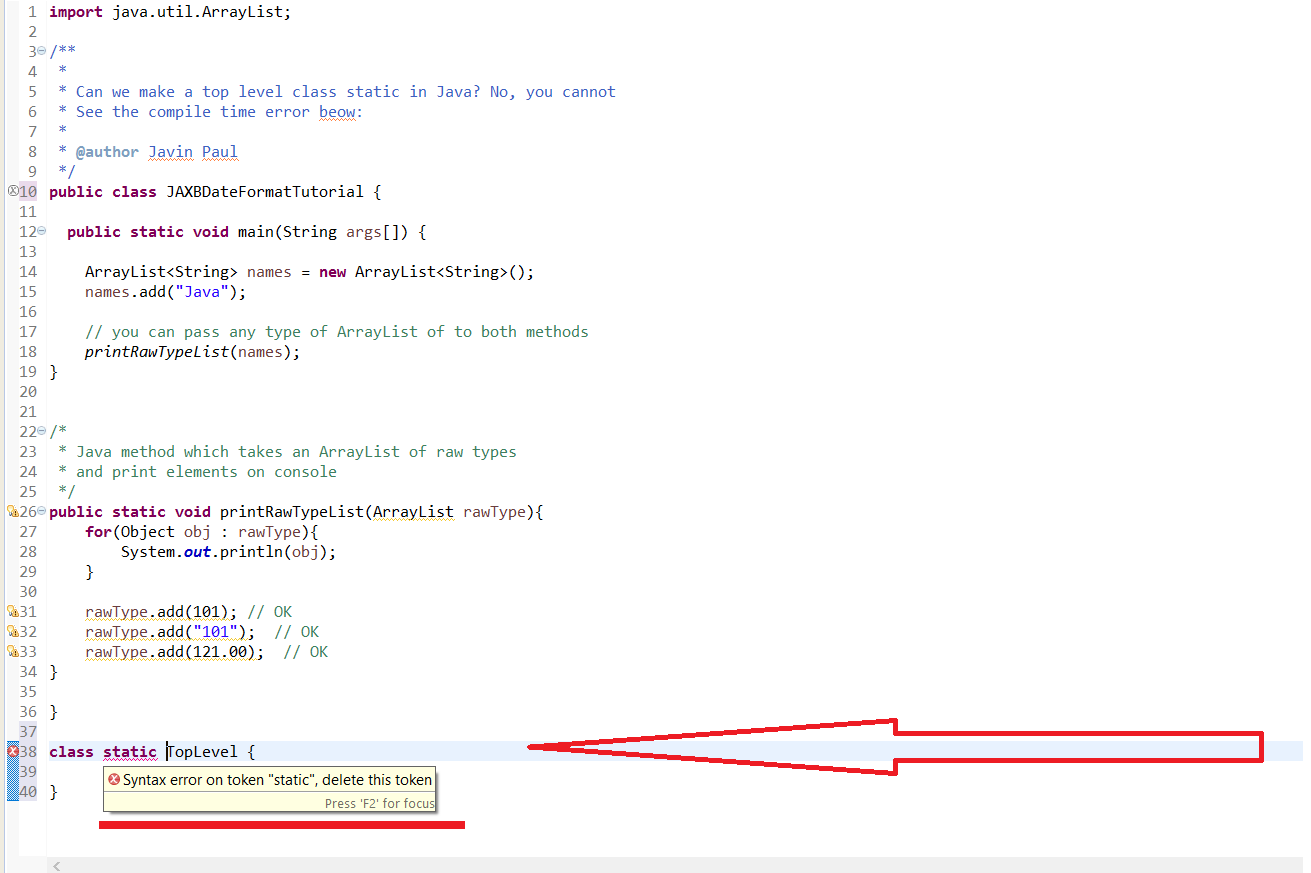 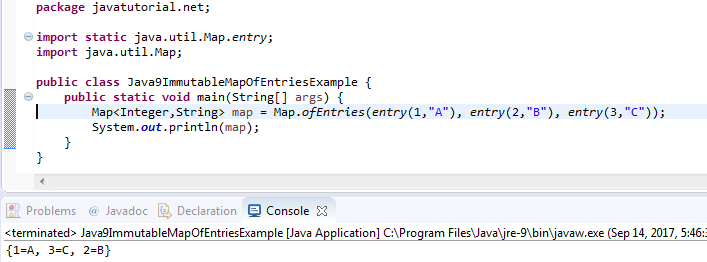
After all, we would expect the drawing of a one to be more similar to other drawings of a one than to a drawing of a zero. When the samples are sorted by label, squareish patterns emerge in the distance matrix. Because a distance matrix of the unsorted samples doesn’t look like much, we also calculate it after sorting the samples by label.įrom trics import pairwise_distancesĭistance_matrix_sorted = pairwise_distances(X_sorted,Īx.imshow(distance_matrix_sorted, 'Greys') Once again we can use scikit-learn to calculate the euclidean distance matrix. The above definition of euclidean distance for two features extends to n features (p 1,p 2,p 2,…,p n). Image credit licensed under CC BY 4.0 by Kmhkmh. Definition of euclidean distance for two features. There are many different distance metrics that make sense but probably the most straightforward one is the euclidean distance. However, even at 728 features, each point is a certain distance apart from every other point. Plotting data with that many features is impossible and that is the whole point of dimensionality reduction. In the actual data, each point is described by 728 features (the pixels). In our t-SNE embedding above, each sample is described by two features. The first step of t-SNE is to calculate the distance matrix. Before we dive into the parameters, we will go through t-SNE step by step and take some looks under the hood of the scikit-learn implementation. Lim = (tsne_result.min()-5, tsne_result.max()+5)Īx.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.0)Ĭonsidering that we did not specify any parameters except n_components, this looks pretty good. # A lot of the stuff here is about making the plot look pretty and not TSNE # Plot the result of our TSNE with the label color coded # We want to get TSNE embedding with 2 dimensions
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
